Missing
- Rotation
- complete R application
Similar to cluster analysis, factor analysis is a method used to identify patterns in data. The difference here is that the groups are not formed from the objects or observations in the data. Instead, single variables are grouped together.
This for example allows us to:
- simplify the data structure by reducing a large number of observed variables to a smaller set of underlying factors
- identify hidden patterns in how variables correlate with each other and uncover latent, underlying dimensions
- create indicators for theoretical constructs
There are two different methods of factor analysis who have slightly different assumptions. The basic idea is, however, to create a correlation matrix of all variables and then group them into a number of (previously determined) factors.
Cluster vs. factor analysis
- Cluster analysis groups objects / cases based on similarities (similar values on a set of variables)
- Factor analysis groups variables based on their correlations
Overview over the two different methods
| Principal Components Analysis | Factor Analysis | |
|---|---|---|
| Variance | completly covered by the factors | Common vs. unique variance |
| Model assumptions | No error / unique factors | Includes unique factors |
| Communality | Always = 1 (all factors are extracted) | Always < 1 |
| Uniqueness | Always = 0 | Always > 0 |
| When to use? | For reducing data into a lower number of dimensions | To find out about underlying latent variables |
Principal Components Analysis (PCA)
Formula:
- : The observed value of unit on variable
- : Factor Loading of variable k on factor
- : Factor score of unit i on factor q
Factor Analysis
Factor analysis assumes that not all of the variance in the data can be traced back to the common factors but that some of it is due other influences (e.g. measurement errors)
Formula:
The formula is the same as for PCA except the last product () which represents the variance which is not accounted for by the common factors.
Relevant concepts
| Concept | Description |
|---|---|
| Factor Loadings | Correlation between a single variable and a factor. |
| Factor Scores | A specific value a specific observation has on a factor (one score per factor per observation) |
| Eigenvalue | Amount of variance explained by a single factor across all variables. |
| Communality | The variance of a single variable accounted for by all the factors. |
| Uniqueness | The variance of a single variable not accounted for by all the factors. |
Hint
- Factor loadings are used to interpret the factors (what variables form a factor together).
- Factor scores show how much of a factor applies to each observation.
Factor loading
The factor loading describes the relationship between a (observed) variable variable and the (unobserved) factor . They are properties of the variables, not the observations.
Meaning: Loadings tell you what the factor represents: A high factor loading means that the variable is strongly associated with, or well represented by, the factor.
Value range: Factor loadings can take values from -1 to 1 (as they are essentially correlations betwen variables and a factor).
In principal component analysis (for z-standardized variables and as long as no skewed rotation was performed), the factor loadings indicate the correlation between a certain variable k and a certain extracted factor.
Squared factor loadings: When the factor loading is squared it indicates the variance of a single variable that is explained by the factor (similarly to the sums of squares in OLS).
Example
Variable Factor 1 Loading Squared Loading Explained variance Math 0.7 0.49 49% Science 0.8 0.64 64% English 0.3 0.09 9% Interpretation:
- Science loads 0.8 and is strongly related to factor 1
- Science loads 0.3 and is weakly related to factor 1
Factor score
The factor score is the estimated score each observation has on one of the factors (i.e. the value of the factor for each case). They are properties of the observations, not of the variables.
Eigenvalue
The Eigenvalue indicates the total variance explained by a factor. It consists of the sum of the squared factor loadings of all variables for a single factor.
- = Number of variables
- = Eigenvalue of factor
- = Factor loading of variable on factor
Because the squared factor loading of each variable can range from 0 to 1, the maximum possible Eigenvalue of each factor corresponds to the number of variables.
Example
The factor analysis contains 10 variables. The theoretical maximum Eigenvalue of any of the factors is 10. An Eigenvalue of 10 would mean that all of the variance of the variables is explained by that single factor.
Communality
The communality is the shared variance on a single variable across all factors (i.e. the sum of the squared factor loadings). IIt shows how much of the variable’s variance is accounted for by the extracted factors.
Formula:
Reading: The communality of the variable consists of the squared factor loadings of variable which are summed up over all the individual factors .
Example
Let‘s take an example variable in a factor analysis with four factors. The loadings of each factor for this specific variable are
-0.07,0.74,0.17, and0.08.The calculation of the communality then looks like this:
This means that 59 % of the variance of that variable is explained by the four factors.
Uniqueness
The uniqueness is the opposite of the communality. It indicates the share of the variance (of a single variable) that is not explained by the factors.
Formula:
Rotation
Application in R
Dataset
The dataset contains observations on countries and variables on different aspects of the political system such as the number of parties, the degree of federalism and so on.
1. Preparing the dataset and creating a correlation matrix
vatter <- import("Vatter2009.dta")
# Data wrangling
vatter_short <- vatter %>%
select(-country, -year) # keep all variables except "country" and "year"
row.names(vatter_short) <- vatter$country
# Correlation matrix with `cor()`:
vatter_cor <- cor(vatter_short)
vatter_cor[upper.tri(vatter_cor)]<- NA # Setting the "upper triangle" as missings for easier visual interpr
View(round(vatter_cor, digits=4))2. Performing a Principal Component Analysis
The command principal performs a PCA. The first argument indicates the correlation table used for the analysis, nfactors indicates the number of factors we want to extract. Here we assume 4 underlying factors. rotation is not applied.
pca_results <- psych::principal(cor(vatter_short),
nfactors=4,
rotate="none")
pca_resultsOutput (split in two):
Principal Components Analysis
Call: psych::principal(r = cor(vatter_short), nfactors = 4, rotate = "none")
Standardized loadings (pattern matrix) based upon correlation matrix
PC1 PC2 PC3 PC4 h2 u2 com
effparties -0.07 0.74 0.17 0.08 0.59 0.41 1.2
execleg 0.25 0.73 -0.23 -0.10 0.66 0.34 1.5
disprop 0.09 -0.76 0.31 0.31 0.78 0.22 1.7
interest -0.41 0.67 -0.31 0.22 0.76 0.24 2.4
centralbank -0.48 0.51 0.02 0.56 0.80 0.20 2.9
federal 0.88 0.10 -0.04 0.08 0.80 0.20 1.0
taxes 0.70 0.26 0.05 -0.44 0.76 0.24 2.0
bicam 0.83 -0.03 0.24 0.34 0.86 0.14 1.5
constitution 0.66 0.33 -0.25 -0.09 0.61 0.39 1.8
judreview 0.65 0.04 -0.09 0.52 0.71 0.29 2.0
cabinets -0.17 0.41 0.73 -0.18 0.75 0.25 1.8
directdem 0.09 0.36 0.75 0.04 0.70 0.30 1.5
PC1 PC2 PC3 PC4
SS loadings 3.33 2.85 1.50 1.10
Proportion Var 0.28 0.24 0.13 0.09
Cumulative Var 0.28 0.52 0.64 0.73
Proportion Explained 0.38 0.32 0.17 0.13
Cumulative Proportion 0.38 0.70 0.87 1.00
Mean item complexity = 1.8
Test of the hypothesis that 4 components are sufficient.
The root mean square of the residuals (RMSR) is 0.09
Fit based upon off diagonal values = 0.92The first part of the output shows the factor loadings for each variable (lines) + each factor (columns)
- E.g. for the variable
effpartiesthe second factor (0.74) can explain a high share of the variance in the variable. To calculate the exact variance the factor loading can be squared u2shows the Uniquenessh2shows the Communalitycomshows complexity (not covered here)
The second part shows information on the factors
SS loadingshows the EigenvaluesProportion Varshows the proportional variance explained by one factor = Eigenvalues (orSS loadingdivided by the number of variablesCumulative Varshows an ongoing addition of the proportional variances from one to another (would go up to 1)
The results can only be display partially using the following commands:
pca_results$loadings
pca_results$uniquenesses3. Creating a scree plot
psych::VSS.scree(cor(vatter_short),
main="Scree plot of eigenvalues by factor")
abline(1,0, col="red")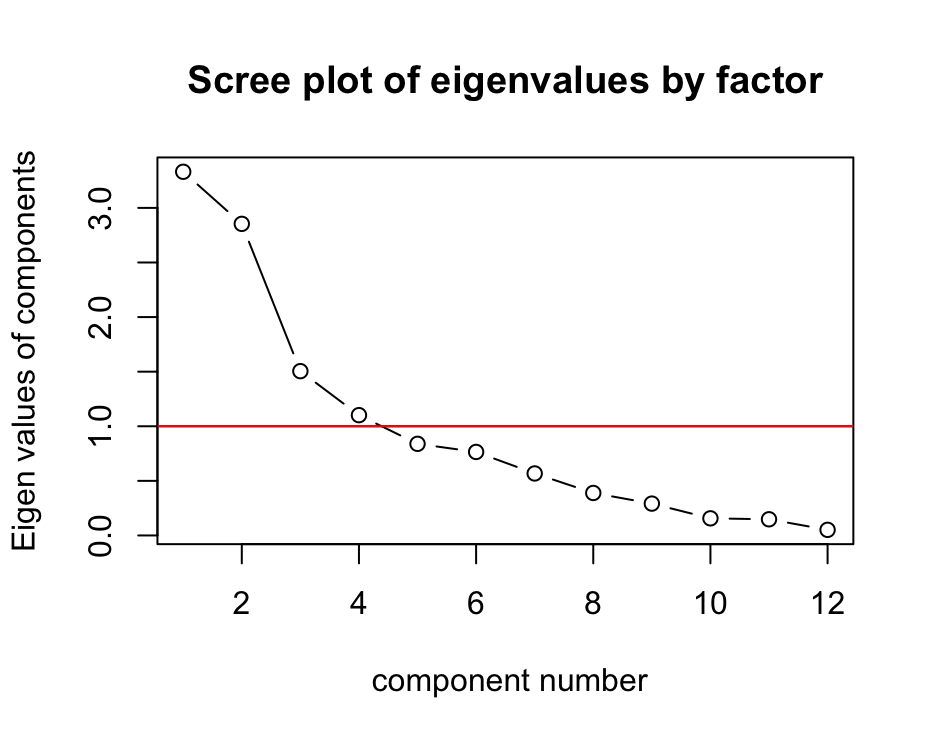 Interpretation:
Interpretation:
- The y-axis shows the eigenvalues of components.
- The x-axis shows number of components.
- The line shows how the eigenvalues drop with an increasing number of components.
According to the Kaiser-Criterion (a rule of thumbs which says to retain factors with eigenvalues that are greater that one should be extracted), the plot shows that four factors should be extracted.
4. Rotation
Goal: Maximising the variance explained by the factors and make sure that each variable only fits to one factor (to the extent possible)
pca_results_rotated <- psych::principal(cor(vatter_short),
nfactors=3,
rotate="Varimax")
pca_results_rotatedVarimax = Orthogonal rotation (preferable)
Promax = Oblique rotation
round(pca_results_rotated$rot.mat, digits=4)Rotation matrix:
[,1] [,2] [,3]
[1,] 0.9862 -0.1568 -0.0525
[2,] 0.1647 0.9036 0.3954
[3,] -0.0145 -0.3986 0.91704. Predicting and interpreting factor scores
factor_scores <- predict(pca_results,vatter_short) PC1 PC2 PC3
Australia 1.56317761 -1.15042788 0.265304376
Austria 0.22205895 0.40163084 -0.752673621
[...]
UK -0.57633764 -2.35926295 0.170078988
USA 2.03517192 -0.38503274 -1.080843363